This package makes it easy to visualize, manipulate, and export phylogenetic trees in R using an interactive viewer/editor. You can insert this tree viewer into Rmd/Quarto documents make your trees accessible online. If you want to use this for web development outside of R, consider the javascript heat-tree package which this package builds upon.
Quick start
The package includes example data sets that are automatically loaded with the package (e.g. weisberg_2020_mlsa), so you can try it out with minimal effort. After installing the packages, simply run the lines below to get an idea of how it works:
See the “Getting Started” page for interactive examples
library(heattree)
heat_tree(
tree = weisberg_2020_mlsa,
metadata = weisberg_2020_metadata,
aesthetics = c(tipLabelColor = 'host_type'),
layout = 'circular')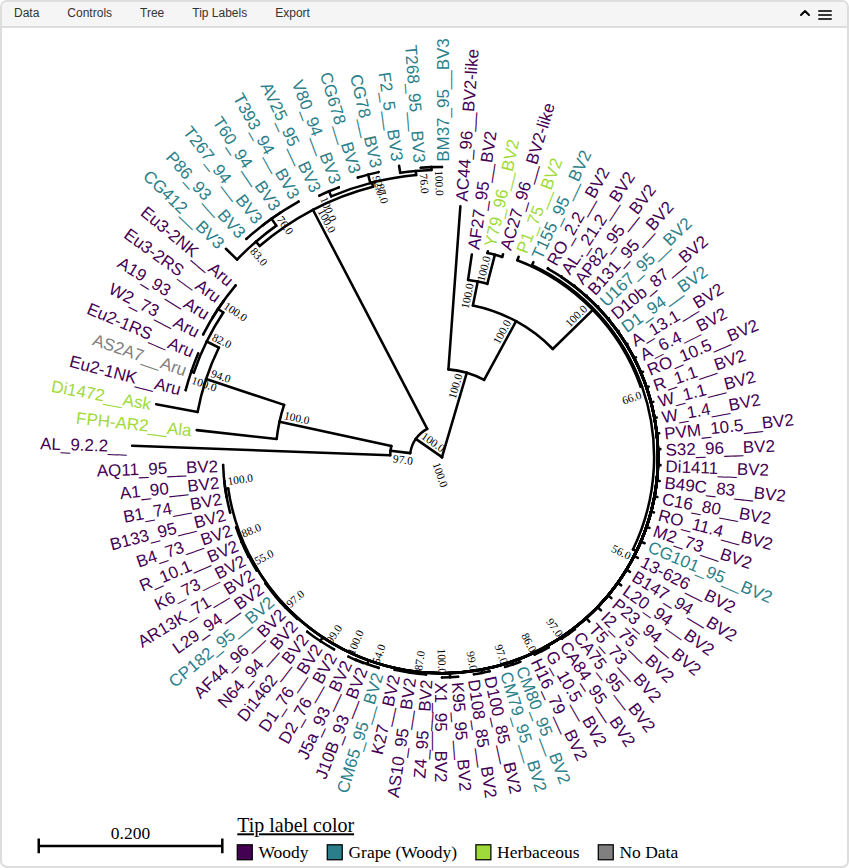
For Python / Jupyter Notebook users
For the python version of this package see this documentation.
For JavaScript users
For the JavaScript library that is the foundation of this package see this documentation.
Contributing and feedback
Contributions and feedback are welcome! Please visit the GitHub repository to report issues or submit pull requests.